Ras activation by NO in breast cancer cells continues to be described to proceed inside a cGMP independ net mechanism and our information exhibiting NO mediated SNO of Ras is steady with this particular prior report. Our locating that NO activation of Ras, by way of SNO results in Ets one activation suggests that other Ras mediated pathways can also activated by NO in human cancer. We propose that the NO/Ets one signaling axis initially described here might advertise sickness progression in other tumors that overexpress NOS2, this kind of as glioma and mela noma, and tumors with impaired SNO metabolism, this kind of as lung and hepatocellular carcinoma. Ets 1 has also been linked to melanoma and lung tumor metas tasis. Moreover, our information displaying that NO effects within a MEK/ERK/Ets one signaling cascade in ER HER2 SKBR3 cells sug gest that high NOS2 expression and NO signaling may possibly induce proliferative and aggressive phenotypes in HER2 breast cancer.
Collectively, these information further strengthen the proposed linkage between NO and Ets 1 signaling and propose that their interaction is actually a significant promoter of tumor metastasis and requires additional investigation. Conclusions In summary, NO signaling effects during the activation in the oncogenic transcription factor our site Ets 1, that’s important for your basal like breast cancer phenotype related with tumor NOS2 expression. This result of NO is mediated by Ras SNO modification and subsequent MEK/ERK signaling to phosphorylate Ets 1. Activation of Ets 1 by NO resulted while in the elevated expression of your basal like markers P cadherin, S100A8, IL 8 and ab crystallin, which mechanistically links two prognostic markers of poor basal like patient survival.
Moreover, NO activation of Ets one resulted in enhanced expression and exercise of proteases essential for tumor metastasis, MMPs and CTSB, and resulted in enhanced cancer cell invasion and prolifera tion. These information imply a molecular mechanism that elu cidates the aggressive selleck basal like phenotype induced by NOS2 and NO signaling and provides a potential thera peutic target for triple negative/basal like breast cancer. Introduction NOTCH activation has been implicated in several malig nancies, notably T cell acute lymphoblastic leukemia, persistent lymphocytic leukemia, glioblastoma, and breast cancer. Overexpression of NOTCH receptors has become implicated in ductal carcinoma in situ and invasive breast cancer, and large ranges of the NOTCH ligand JAG1 appear to predict a poor total sur vival.
High NOTCH1 receptor ranges happen to be linked with basal like, triple detrimental breast cancer, and NOTCH1 levels correlate with abbreviated survival. Additional not long ago, silencing of Lunatic Fringe, the glycosylase that regulates NOTCH1 ligand activity, continues to be observed in individuals with basal like breast cancer, and increased ranges of intracellular NOTCH1 are detected in these patients cells.
Monthly Archives: June 2014
The sequences for Rac1 siRNAs are Cells had been transfected wi
The sequences for Rac1 siRNAs are Cells were transfected with siRNAs at 100 nM by using DharmaFECT1 siRNA transfection reagent, according to the suppliers instruction. For experiments involving both siRNA transfection and IR publicity, transfected cells have been initially incubated for the indicated times and after that exposed to IR. Adenoviral vectors and adenoviral infections Recombinant adenovirus N17Rac1 and control adenovirus dl312 had been kindly professional vided by Dr. Toren Finkel. In Ad. N17Rac1, the Rac1 cDNA has a Ser to Asp substitution at position 17 and functions as a dominant negative mutant. Log phase MCF seven cells were infected at 50 PFU/cell with either Ad. N17Rac1 or Ad. Handle for 24 hours prior to exposure to IR, as described previously.
For scientific studies involving cell cycle analysis, the cells have been incu bated for more 24 hours just after IR and analyzed for DNA content material with movement cytometry. For studies involving mitotic cell evaluation, the irradiated cells were incubated a fantastic read for 2 hours and analyzed for cells containing each 4N DNA material and histone H3 Ser10 phosphor ylation. DAPI staining Apoptosis was assessed with four,6 diamidino two phenylin dole staining, as described previously. Apoptotic cells had been recognized by condensation and fragmentation of nuclei. The percentage of viable cells was calculated since the ratio of live cells to total cells counted. No less than 800 cells had been counted per sample. Cell survival assay Cell survival assays have been performed as described pre viously. In brief, log phase increasing cells have been exposed to IR in the doses indicated, incubated for seven days, and visualized for viable cells by staining with crystal violet.
For experiments involving treatment method with the two NSC23766 and IR, ZSTK474 cells have been preincubated for 1 hour with one hundred uM NSC23766, exposed to IR, and incubated for an addi tional three hrs soon after IR. The cells were washed and incu bated in normal growth medium for seven days before analysis. The obtained sample dishes had been scanned on an EPSON Perfection 4490PHOTO scanner, along with the level of cells remaining within the culture dish was quantified through the use of the ImageJ analytic program. Clonogenic assay Clonogenic assay was performed as described previously. In quick, inside the presence or absence of one hundred uM NSC23766, MCF seven cells had been exposed to IR in the doses indicated and incubated for three hrs right after IR.
The cells had been then rinsed with DMEM, reseeded at the cell num ber indicated in duplicate, and incubated for 10 to 14 days right up until colonies formed. The colonies have been visualized with crystal violet staining and quantified by using Ima geJ application, as described previously. Success IR publicity induces G2/M arrest and Rac1 GTPase activation in MCF seven breast cancer cells To research the mechanisms regulating G2/M cell cycle checkpoint response just after IR exposure, log phase expand ing MCF 7 cells have been exposed to IR in the indicated doses and analyzed for DNA written content at eight, sixteen, and 24 hrs just after IR.
For that reason, GMP compatible recombinant proteins for large sc
Thus, GMP compatible recombinant proteins for significant scale pilot research or preclinical trials need to be developed. The time of application of a tolDC vaccine to pSS sufferers is yet another challenging difficulty. Ordinarily, it requires a number of many years to set up a diagnosis and often at this time the injury to salivary glands is irreversible. Then again, although some scientific studies have observed associations between the degree of destruction and loss of perform, with the patient degree there’s not automatically such a correlation. Conclusions In summary, this is actually the to start with examine demonstrating the suc cessful generation of Ro/La loaded tolDC from sufferers with pSS capable to effectively induce antigen specific sup pressive immune reactions.
Since the described protocol can conveniently selleck be adjusted to get GMP compatible, we believe that tolDC might be a possibly favourable treatment method alternative for individuals with pSS with identified antibody specificity. The traditional Warburg result versus oxidative mitochondrial metabolic process The Warburg eect, also referred to as aerobic glycolysis, is dened because the propensity of cancer cells to consider up substantial amounts of glucose and to secrete lactate while in the presence of oxygen. Warburgs unique function indicated that when glucose uptake and lactate manufacturing are tremendously elevated, a cancer cells fee of mitochondrial respiration is similar to that of regular cells. He, however, described it as a respiratory impairment as a result of undeniable fact that, in cancer cells, mitochondrial respiration is smaller sized, relative to their glycolytic energy, but not smaller sized relative to usual cells.
He recognized that oxygen consumption is not diminished in tumor cells, but that respiration is disturbed simply because selleck chemical GSK1210151A glycolysis persists in the presence of oxygen. However, the perception of his unique ndings was simplied above the years, and most subse quent papers validated that cancer cells undergo aerobic glycolysis and produce lactate, but did not measure mitochondrial respiration, and just presumed decreased tricarboxylic acid cycle action and reduced oxidative phosphorylation. It is certainly effectively docu mented that, being a consequence of intra tumoral hypoxia, the hypoxia inducible aspect one pathway is activated in lots of tumors cells, resulting in the direct up regulation of lactate dehydrogenase and enhanced glucose consumption. For up to date reviews to the Warburg eect, the reader is encouraged to refer to the following papers. Having said that, new ndings compel us to reconsider the present model of cancer cell metabolism. To start with, not all tumors are associated with increased aerobic glycolysis, and in reality it really is now clear that cancer cells employ both glycolysis and oxidative phosphorylation to satisfy their metabolic desires.
Gene Set Enrichment Evaluation was carried out using gene sets de
Gene Set Enrichment Examination was carried out using gene sets derived from published literature. So as to stay clear of false positives due to multiple testing in GSEA, the false discovery rate was utilised to alter the P value to give the Q value. A Q worth of 0. 05 is statistically substantial. X box binding protein mRNA splicing assay XBP one mRNA was amplified from 50 ng cDNA utilizing 0. six uM primers, 250 mM MgCl2, and 0. 25 U of Less complicated Red Taq DNA polymerase inside a last volume of 25 uL, at an anneal ing temperature of 66 C for 35 cycles. Forward primer. PCR goods have been digested with PstI and separated on the 3% agarose gel. A 448 base pair amplicon signifies spliced XBP one. Protein synthesis Protein synthesis was established following 92 hrs of gene silencing.
Cells had been washed twice in PBS then incubated for four hours in cysteine/methionine kinase inhibitor SB 203580 cost-free media containing 0. 5% bovine serum albumin, glutamine and 10 uCi of 35S Express Protein Labelling Combine, from the presence of both ethanol or four OHT, then lysed in RIPA buffer. Soluble proteins were precipitated from cell lysates with 25% ultimate concentration of trichloracetic acid and 10 ug BSA. Precipitates have been centrifuged, washed twice in 10% TCA and twice in ethanol, just before scintillation counting. Information were normalized applying complete protein con tent established by sulforhodamine B assay from parallel cultures. Determination of ROS levels Cells have been incubated with three uM CM H2DCFDA for thirty minutes or with 2. five. uM MitoSOX for 15 minutes at 37 C, trypsini zed and washed twice with PBS, stained with DAPI and analyzed on a LSRII SORP movement cytometer.
Examination of cellular respiration Experiments had been carried out inside a 96 effectively format using a Seahorse Bioscience XF96 Extracellular Flux Analyser in Seahorse Bioscience TGX221 assay medium supplemented with one mM sodium pyruvate and ten mM Glucose and pH was adjusted to seven. 4. All through the experiment, 1. 264 uM oligo mycin A, 0. four uM FCCP, along with a mix of 1 uM rotenone and one uM antimycin A had been injected. Oxygen consumption costs were measured more than time and normalized to complete protein con tent established by sulforhodamine B staining. Lipid evaluation by mass spectrometry Lipids were extracted making use of a methanol/chloroform extrac tion process and quantified by Liquid chromatography mass spectrometry analysis on the Shimadzu IT TOF LC/MS/MS program. Correct mass and tandem MS have been used for molecular species identifi cation and quantification.
The identity of lipids was further confirmed by reference to appropriate lipid stan dards. A detailed description of your method is presented inside the More file one supplemental information and facts. Cell viability assay Caspase 3/7 activity was measured using Caspase three sub strate IX, fluorogenic, Cells had been fixed with trichloroacetic acid and normalized to complete protein articles  determined by sulforhodamine B staining.
determined by sulforhodamine B staining.
Coefficients b are sought iteratively in optimum probability esti
Coefficients b are sought iteratively in maximum likelihood estimation. Likelihood displays the estimated probabilities of all N genes belonging to their real class, and thus gives a measure for model eva luation, where yi,c 1 if yi is of class c and 0 otherwise, as well as the probability of gene class romantic relationship is computed as microarrays by Zhu et al. The data were even further professional cessed with in vivo nucleosome positioning measurements to distinguish binding online websites where decrease nucleosome occupancy displays open chromatin framework. Our dataset of 285 regulators includes 128,656 signifi cant associations concerning regulators and target genes. Maximising the log probability l prospects to optimum regression coefficients B and the corresponding likeli hood worth , Statistically reasoned cutoffs render our dataset sparse, it comprises higher self confidence signals to 7.
2% of approxi mately 1. 8 million probable TF gene pairs. The dataset consists of 107 TF target sets with knockout data, sixteen TFs with TFBS predictions and 162 TFs with both varieties of evidence. The majority of all gene regulator associations Right here we implemented a statistical check to assess the pro cess specificity of a provided TF by evaluating two top article multino mial regression designs. The null model H0, g b0 is surely an intercept only model the place system particular genes are predicted solely based on their frequency during the total dataset. The option model H1, g b0 bkXk is really a univariate model during which TF targets are also regarded as predictors of course of action genes.
We make use of the likeli hood ratio check together with the chi square distribution to compare the likelihoods on the two designs, and Vicriviroc come to a decision if incorporating TF information and facts substantially improves match to data offered its more complexity, as the place ? corresponds to degrees of freedom and displays number of model parameters. To predict all reg ulators to a procedure of interest, we test all TFs indepen dently, right for multiple testing and get TFs with significant chi square p values. In summary, m,Explorer makes use of the multinomial regression framework to associate practice genes with TF regulatory targets from TFBS maps, gene expression patterns and nucleosome positioning information. Our technique finds candidate TFs whose targets are particularly informative of method genes, and thus may well regulate their expression.
Yeast TF dataset with perturbation targets, DNA binding online websites and nucleosome positioning We made use of m,Explorer to research transcriptional regulation and TF function in yeast, as it has the widest assortment of relevant genome wide evidence. Initially we compiled a data set of 285 regulators that includes carefully picked target genes for almost all yeast TFs from microarrays, DNA binding assays and nucleosome positioning measurements. Statistically sizeable target genes from regulator deletion experiments originate from our latest reanalysis of an earlier research.
Array intensities judged as drastically improved were chosen by t
Array intensities judged as considerably elevated were chosen by two crite ria, p 0. 005, and fold transform one. 4. At least half from the genes also had a constructive B value. The double criteria identified 288 gene promoters, which are listed in Table S1 in Further data file 2. The many information files have already been submitted to. Confirmation on the differential expression of UV induced genes working with bioinformatics criteria A number of observations indicate the substantial alterations observed here accurately reflect differential precipitation and array binding. 1st, for the 283 genes that exhibited signifi cantly altered hybridization following UV irradiation, 112/ 283 have ideal Egr1 consensus websites within their pro moter sequences. A different 53 genes have probable EBSs whereas the frequency of EBSs inside a set of 200 random sequences was only 23%.
As a result, the professional moters reported as bound by Egr1 certainly consist of a signifi cant increase inside the frequency of EBSs. Secondly, at the least 43/ 283 genes are acknowledged to get UV responsive from other studies. A third indication comes from the identification of 24/283 drastically bound genes as EGFR linked genes. These genes had been recognized by Pathway studio five. 0, which compiles citations selleck inhibitor indicating that expression of these genes is linked with EGFR action and/or expres sion. To evaluate this frequency, a set of 1,000 genes was examined in Pathway studio 5. 0 making use of the same query, which yielded only 26 genes related to EGFR. We examined the functional nature with the identified genes utilizing plan assisted literature surveys like Ariadne and Ingenuity.
Numerous functional groups of genes have been apparent. These involve regulators of apoptosis for instance Bcl G, BLK, CASP7, BBC3 as well as TNFSF5, TNFSF6 and TNFSF19L, which belong towards the tumor necrosis issue loved ones. Genes encoding the DNA repair enzymes NT5E, NME1 and NME2, cytokines, for example IL1R1, IL15 and IL18R1, the cell cycle regulators CDK8, CDKN1b/ p27, PAK6 and selleck chemicals SKP1a plus the transcription regulators Ets2, Egr2, POU4F1, SOX11, EN1 and HSF4 had been all amongst those containing significantly detected promoters. Genes for instance BBC3, PTPN13, MAX, MAP3K7 and MAP2K1 and 38 many others, have been previously documented as UV responsive genes. Experimental validation of hybridization intensities Traditional ChIP was performed to verify the results of ChIP on chip experiments making use of a set of 25 representative genes.
Primers had been created close to the putative EBS around the target promoters and these had been utilised for qRT PCR amplifica tion of your sequences from your ChIP captured chromatin. The qRT PCR success demonstrate that in 23/25 genes, UV treatment method led to increased PCR yields of 1. 4 to eight fold when compared to control cells. In contrast, tiny or no DNA enrichment was observed for all 25 primer sets when applied to precipitates ready using handle IgG serum.
If that is the situation, a primer pair will match to two differ
If this is often the situation, a primer pair will match to two differ ent sequences. Of the 173 SSR markers existing within the N. acuminata genetic map, 128 of them could be mapped to your N. sylvestris genome assembly. This number may be the sum of the 75 SSRs of the N. acuminata map uncovered while in the N. sylvestris selleck chemical assembly, the 50 SSRs of the N. acuminata map located from the N. sylvestris and N. tomentosiformis assemblies, the single SSR of the N. acuminata and N. tomentosiformis maps observed in the N. sylvestris assembly, plus the two SSRs in the N. acuminata and N. tomentosiformis maps found in the N. sylvestris and N. tomentosiformis assemblies. Similarly, within the 221 SSR markers present within the N. tomentosiformis genetic map, 173 may be mapped to the N. tomentosiformis gen ome assembly.
Moreover, 706 SSR markers not existing to the present genetic maps can be mapped to the N. sylvestris genome assembly, 605 mapped for the N. tomentosiformis genome assembly, MK-2048 and 174 mapped to each. Of your 134 COSII markers existing during the N. acumi nata genetic map, 45 could possibly be mapped on the N. sylvestris genome assembly. Similarly, of the 262 COSII markers inside the N. tomentosiformis genetic map, 81 can be mapped on the N. tomentosiformis genome assembly. Employing the same approach, 736 in the 879 COSII markers within the expen2000 tomato genetic map could possibly be found, 718 of them mapped to your expected chromo some. Also, 68 COSII markers not existing for the existing genetic maps might be mapped towards the N. sylves tris genome assembly, 78 mapped towards the N. tomentosi formis genome assembly, and 226 mapped to the two.
The very low numbers of COSII markers that can be mapped for the N. sylvestris and N. tomentosiformis assemblies, despite the great benefits that were obtained utilizing the identical process on the tomato map, could possibly be because of the present 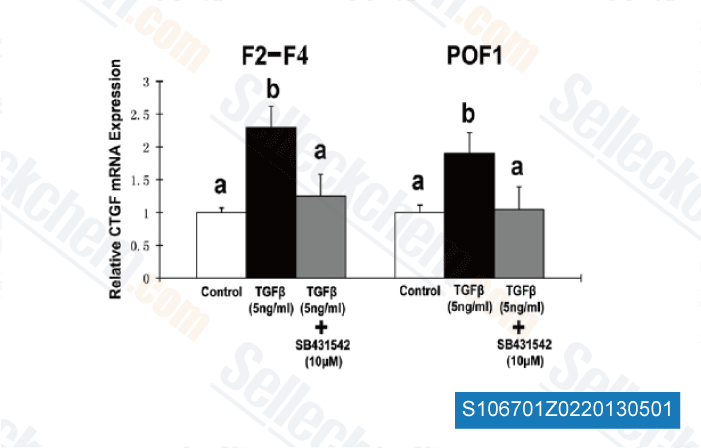 fragmented state in the assemblies, or as the COSII marker primers are certainly not adapted for Nicotiana species. Transcriptome assembly The amount of reads obtained for each in the tissue specific samples from both species is outlined in Addi tional file 9. Tissue precise assemblies were produced for the 3 samples by mapping the reads for the reference genomes implementing the Bowtie2/ Tophat2 pipeline. The length distributions in the assembled transcripts are summarized in table three. Also, a reference transcriptome for every species was made by merging the three person tissue exact assemblies. We also applied a de novo assembly system to produce an assembly that potentially has tran scripts missing through the mapping assembly as a result of the absence of selected genes through the current reference gen
fragmented state in the assemblies, or as the COSII marker primers are certainly not adapted for Nicotiana species. Transcriptome assembly The amount of reads obtained for each in the tissue specific samples from both species is outlined in Addi tional file 9. Tissue precise assemblies were produced for the 3 samples by mapping the reads for the reference genomes implementing the Bowtie2/ Tophat2 pipeline. The length distributions in the assembled transcripts are summarized in table three. Also, a reference transcriptome for every species was made by merging the three person tissue exact assemblies. We also applied a de novo assembly system to produce an assembly that potentially has tran scripts missing through the mapping assembly as a result of the absence of selected genes through the current reference gen
Remedy with PD153035 inhibited Egr1 expression by roughly 85% and
Therapy with PD153035 inhibited Egr1 expression by approximately 85% and suramin inhibited Egr1 expression by somewhere around 80%. In addition, our ChIP on chip benefits showed that EGFR expression was sup pressed by Egr1 upon UV irradiation and greater by threefold when the cells were irradiated following silencing Egr1 expression. The result signifies that Egr1 promoter binding is specifically linked with decreased transcription of EGFR, suggesting the presence of a unfavorable feedback loop controlling EGFR expression by Egr1. Egr1 above expression right after UV irradiation leads to growth inhibition and apoptosis UV stimulation promotes apoptosis within a selection of cell kinds. We therefore examined the development and survival properties of larly, in M12 cells we observed that ERK1/2 inhibitors block M12 cells following UV stimulation by direct proliferation measurements more than three days.
Untreated M12 cells in conventional medium grew rapidly selelck kinase inhibitor to substantial density whereas cells treated by UV irradiation have been drastically retarded in development, which was apparent within 24 h. By 24 h numerous detached and floating cells and extracellular debris had been apparent, sug gesting apoptosis in these cells. A Poly ribose polymer ase assay unveiled a higher proportion of PARP degradation, indicating apoptosis, whereas no degradation was obvious in untreated cells. Cell numbers have been diminished 25 fold compared to control cells at 72 h right after treatment method. These success indicate that EGFR activation leads to apoptosis in M12 prostate cells. To test whether apoptosis of M12 cells was Egr1 dependent in vivo, M12 cells have been taken care of with siEgr1 to silence Egr1 expression for 48 h followed by UV C.
Egr1 mRNA and pro tein expression was proficiently silenced by this treatment. Cells have been collected 24 h later on as well as the PARP assay demonstrated that cells underwent lowered apoptosis while in the absence of Egr1, obviously displaying that Egr1 is definitely an essential mediator selleck RAF265 of UV C induced apoptosis. These final results verify the role of Egr1 as a mediator in the apoptosis response. Discussion Egr1 binds a substantial spectrum of promoters that lead to transcriptional regulation We examined the part of Egr1 in UV irradiated tumorigenic human M12 prostate cancer cells. Our information show that Egr1 binds to a surprisingly substantial quantity of promoters of an array containing around ten,012 special proximal promoter sequences. Numerous of our observations propose that Egr1 promoter binding contributes to your regula tion of gene expression in UV handled cells. 1st, 5. 2% of the drastically bound genes are known to interact with Egr1 and most of them are identified to be regu lated by Egr1. For examination ple, DMRT1 and EGFR are both shown to get direct targets of Egr1 and Egr1 binds to their promoters.
Other observed AEs had been also constant with people of MK 2206
Other observed AEs were also steady with those of MK 2206 single agent treatment. The blend of MK 2206 and trastuzumab also demonstrated preliminary evidence of therapeutic efficacy in sufferers with HER2 breast cancer or gastroesophageal cancer, having a clinical benefit response price of approxi mately 24% as well as a median time for you to progression of 72 days. One patient with metastatic breast cancer, whose disorder progressed about the proper chest wall all over the previous mastectomy scar whilst on servicing therapy with tras tuzumab, accomplished CR following combination therapy with MK 2206. Her erythematous chest wall skin lesion showed a dramatic improvement soon after obtaining two cycles of study remedy and by six months the skin lesion had wholly resolved.
There was a single extra patient with breast can cer handled for more than a yr encountering a complete reduction in tumor size of 68% selleck chemicals who was confirmed as getting PR. 5 more patients had SD for greater than 4 months. These preliminary efficacy benefits recommend the combination of MK 2206 with trastuzumab may perhaps present patients an efficient salvage routine following progression on trastuzumab, or might avoid or delay clinical resistance if applied earlier while in the illness. The efficacy observed within this phase 1 study supports the hypothesis that a mechanism of resistance to trastu zumab could possibly be mediated by activation with the PI3K/AKT pathway in vivo. The mechanisms as a result of which the PI3K/AKT pathway could be activated in trastuzumab refractory HER2 tumors is now unknown.
Major candidates involve activating mutations from the PIK3CA gene or deletion or mutations in PTEN, an inhibitor with the PI3K/AKT pathway. We collected circulating nucleic acid to discover this likelihood, primarily based on reviews that cor connected findings in circulating nucleic acid Tubastatin A with DNA from tumor specimens. Only 3 individuals have been uncovered to get mutations in the PIK3CA gene in circulat ing DNA and none had notably lengthy SD or response to remedy. No PIK3CA mutation was detected from the circulating nucleic acid samples from sufferers who responded to treatment method. Research have estimated that in between 13 and 31% of HER2 breast cancers harbor mutations in PIK3CA. Success of PIK3CA mutation standing from circulating DNA within this examine are at the lower limit of those estimations. Among the limitations of this evaluation is the fact that our PIK3CA mutation evaluation was restricted to circulating DNA evaluation.
Tumor biopsies for biomarker evaluation just before treat ment were not mandated and intratumor heterogeneity in PIK3CA mutation standing or limitations of detection inherent to circulating DNA mutational evaluation could possibly be responsible for that reduce than expected PIK3CA muta tional frequency observed. The probability for that reason re mains that tumor samples at major or metastatic web-sites may possibly demonstrate mutations that don’t seem in circulating nucleic acid.
All cDNA tem plates were amplified by using a pair of popular PCR
All cDNA tem plates have been amplified with a pair of widespread PCR primers. The primer on the strand complementary for the array was fluorescently labeled for subsequent hybridization on the arrays. Validation on the picked miRNAs, proven to be regulated by Illumina miRNA microarray, was carried out by RT PCR. QRT PCR was carried out using the RT2 ProfilerTM Human miFinder miRNA PCR Array from SuperArray. RT2 Profiler PCR Arrays are designed for relative quantitative QRT PCR based on SYBR Green detection and carried out on the one particular sample/one plate 96 well format, utilizing primers to get a preset listing of 88 most abundantly expressed and best characterized micro RNA sequences. In brief, miRNA was converted to cDNA by means of a universal tailing and reverse transcription response. CDNA volumes were adjusted to 2.
5 ml with SuperArray RT2 Serious ABT-737 852808-04-9 Time SYBR Green/ROX PCR 2X Master Combine and 25 ul of cDNA combine was extra to all wells. The PCR plate was sealed and spun at 1500 rpm X four min. True time PCR was carried out on an Utilized Biosystem 7300 Real Time PCR Technique. ABI instrument settings integrated setting reporter dye as SYBR, passive reference is ROX, Delete UNG Activation, and include Dissociation Stage. To correlate differentially expressed miRNAs and their regulated genes, we applied differentially regulated and picked miRNAs towards an established miRNA database for pre dicted target genes. MicroRNA data was also analyzed by the use of Ingenuity Pathway Analysis. Pathway enrichments have been calculated employing the NIAID DAVID functional enrichment device.
Statistical examination Preliminary examination Camostat Mesilate of the scanned information was carried out employing Illumina BeadStudio application which returns single in tensity data values/miRNA following the computation of a trimmed imply average for each probe type represented by a variable quantity of bead probes/gene to the array. Data was globally normalized by scaling just about every array to a prevalent me dian worth, and significant improvements in gene expression be tween class pairs had been calculated using the Student t test. Substantial gene lists had been calculated by picking out genes which content a significance threshold criteria of t test p values much less than or equal to 0. 05 plus a fold transform two or greater. Relative miRNA expression derived from QRT PCR was calculated by utilizing the 2 Ct technique, during which Ct signifies cycle threshold, the fractional cycle amount where the fluorescent signal reaches detection threshold.
The normalized Ct value of each sample is calculated using an endogenous control compact molecular bodyweight RNA. Fold transform values are presented as common fold transform two for genes in taken care of relative to control samples. The criteria of significance used for that RT PCR final results have been the identical as utilized for the Illumina miRNA arrays. Success Demographic traits Demographic  characteristics for all review participants were equivalent in all treatment groups.
characteristics for all review participants were equivalent in all treatment groups.
